Conformers:
ABBIImiBZICOPNSYNN
Click the Summary/Torsions/Similar/... tabs for more details.
SOLUTION NMR STRUCTURE OF A SUBSTRATE FOR THE ARCHAEAL PRE-TRNA SPLICING ENDONUCLEASES: THE BULGE-HELIX-BULGE MOTIF, MINIMIZED AVERAGE STRUCTURE
Click a row in table or a step in viewer for analysis of results. Click column headers to sort data.
Steps with non-standard or missing atoms have not been assigned, description of conformers is defined in the table.
Results of the assignment of 37 detected steps in 1 model(s), can be also downloaded as csv or json file. Average confal 0, percentile 0.
Click a row in table or a step in viewer for analysis of results. Click column headers to sort data.
Define restraints for steps within Å cartesian RMSD, global sigma scaling factor or use per step sigma scaling.
| step number | Step name | CANA | NtC | confal | rmsd |
|---|---|---|---|---|---|
| 01 | 2a9l_A_G1_G2 | AAA | AA00 | 54 | 0.22 |
| 02 | 2a9l_A_G2_G3 | AAA | AA00 | 54 | 0.23 |
| 03 | 2a9l_A_G3_U4 | AAA | AA00 | 70 | 0.18 |
| 04 | 2a9l_A_U4_G5 | AAA | AA00 | 54 | 0.25 |
| 05 | 2a9l_A_G5_A6 | NAN | NANT | 0 | 1.07 |
| 06 | 2a9l_A_A6_C7 | NAN | NANT | 0 | 1.16 |
| 07 | 2a9l_A_C7_U8 | AAA | AA00 | 90 | 0.17 |
| 08 | 2a9l_A_U8_C9 | AAA | AA00 | 62 | 0.28 |
| 09 | 2a9l_A_C9_C10 | AAA | AA00 | 72 | 0.28 |
| 10 | 2a9l_A_C10_A11 | NAN | NANT | 0 | 0.70 |
| 11 | 2a9l_A_A11_G12 | NAN | NANT | 0 | 1.32 |
| 12 | 2a9l_A_G12_A13 | NAN | NANT | 0 | 1.46 |
| 13 | 2a9l_A_A13_G14 | NAN | NANT | 0 | 0.56 |
| 14 | 2a9l_A_G14_G15 | AAA | AA00 | 78 | 0.20 |
| 15 | 2a9l_A_G15_U16 | AAA | AA00 | 71 | 0.25 |
| 16 | 2a9l_A_U16_C17 | AAA | AA00 | 61 | 0.28 |
| 17 | 2a9l_A_C17_G18 | AAA | AA08 | 46 | 0.49 |
| 18 | 2a9l_A_G18_A19 | NAN | NANT | 0 | 0.81 |
| 19 | 2a9l_A_A19_G20 | AAA | AA09 | 37 | 0.30 |
| 20 | 2a9l_A_G20_A21 | AAA | AA02 | 77 | 0.22 |
| 21 | 2a9l_A_A21_G22 | NAN | NANT | 0 | 0.66 |
| 22 | 2a9l_A_G22_A23 | AAA | AA00 | 88 | 0.10 |
| 23 | 2a9l_A_A23_C24 | AAA | AA00 | 73 | 0.16 |
| 24 | 2a9l_A_C24_C25 | A-B | AB05 | 17 | 0.33 |
| 25 | 2a9l_A_C25_G26 | NAN | NANT | 0 | 1.60 |
| 26 | 2a9l_A_G26_G27 | AAu | AA12 | 24 | 0.54 |
| 27 | 2a9l_A_G27_A28 | AAA | AA00 | 25 | 0.41 |
| 28 | 2a9l_A_A28_G29 | AAA | AA00 | 55 | 0.29 |
| 29 | 2a9l_A_G29_A30 | AAA | AA00 | 51 | 0.29 |
| 30 | 2a9l_A_A30_U31 | NAN | NANT | 0 | 0.82 |
| 31 | 2a9l_A_U31_A32 | NAN | NANT | 0 | 0.95 |
| 32 | 2a9l_A_A32_U33 | NAN | NANT | 0 | 0.38 |
| 33 | 2a9l_A_U33_C34 | AAA | AA00 | 66 | 0.26 |
| 34 | 2a9l_A_C34_A35 | AAA | AA00 | 44 | 0.34 |
| 35 | 2a9l_A_A35_C36 | AAA | AA00 | 52 | 0.22 |
| 36 | 2a9l_A_C36_C37 | AAA | AA00 | 57 | 0.26 |
| 37 | 2a9l_A_C37_C38 | AAA | AA00 | 49 | 0.31 |
Steps with non-standard or missing atoms have not been assigned, description of conformers is defined in the table.
|
step: Table of conformers |
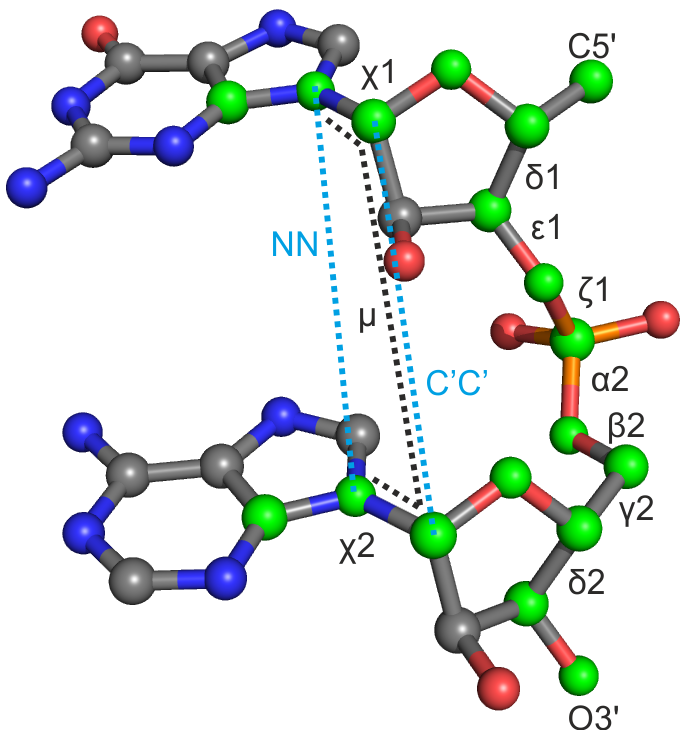
| δ1 | ε1 | ζ1 | α2 | β2 | γ2 | δ2 | χ1 | χ2 | μ | NN | CC | |
| step_torsions | ||||||||||||
| ntC_average | ||||||||||||
| Δ torsions | ||||||||||||
| torsion scores |
| comments | step confal = NaN cartesian RMSD = NaN Å pseudorotation: NA details: NA |
Similarity of step to class averages
Download
Results as csv or json file.Best NtC fitted to input structure.
Restraints
Restraints file for REFMAC (steps with RMSD <= 0.5Å)Commands file for MMB (steps with RMSD <= 0.5Å)
Restraints file for Phenix (steps with RMSD <= 0.5Å) [?]
Definitions
Average parameters (csv), esd values (csv), and Cartesian coordinates of conformers.Download the papers
Description of DNATCO server:Černý et al., Nucleic Acids Research, 44, W284 (2016).
Definition of conformers:
Schneider et al., Acta Cryst D, 74, 52-64 (2018).
Example of application:
Schneider et al., Genes, 8(10), 278, (2017).
Browse conformers
Show
| Color by conformation group (pyramids) | |
|---|---|
| group | |||||||||
| visible |
| Color by NtC (balls) |
|---|
| A | ||||
|---|---|---|---|---|
| A-B | ||||
|---|---|---|---|---|
| B-A | ||||
|---|---|---|---|---|
| B | ||||
|---|---|---|---|---|
| IC | ||||
|---|---|---|---|---|
| OPN | ||||
|---|---|---|---|---|
| Z | ||||
|---|---|---|---|---|
| N | ||||
|---|---|---|---|---|